Deep learning
for spectral science
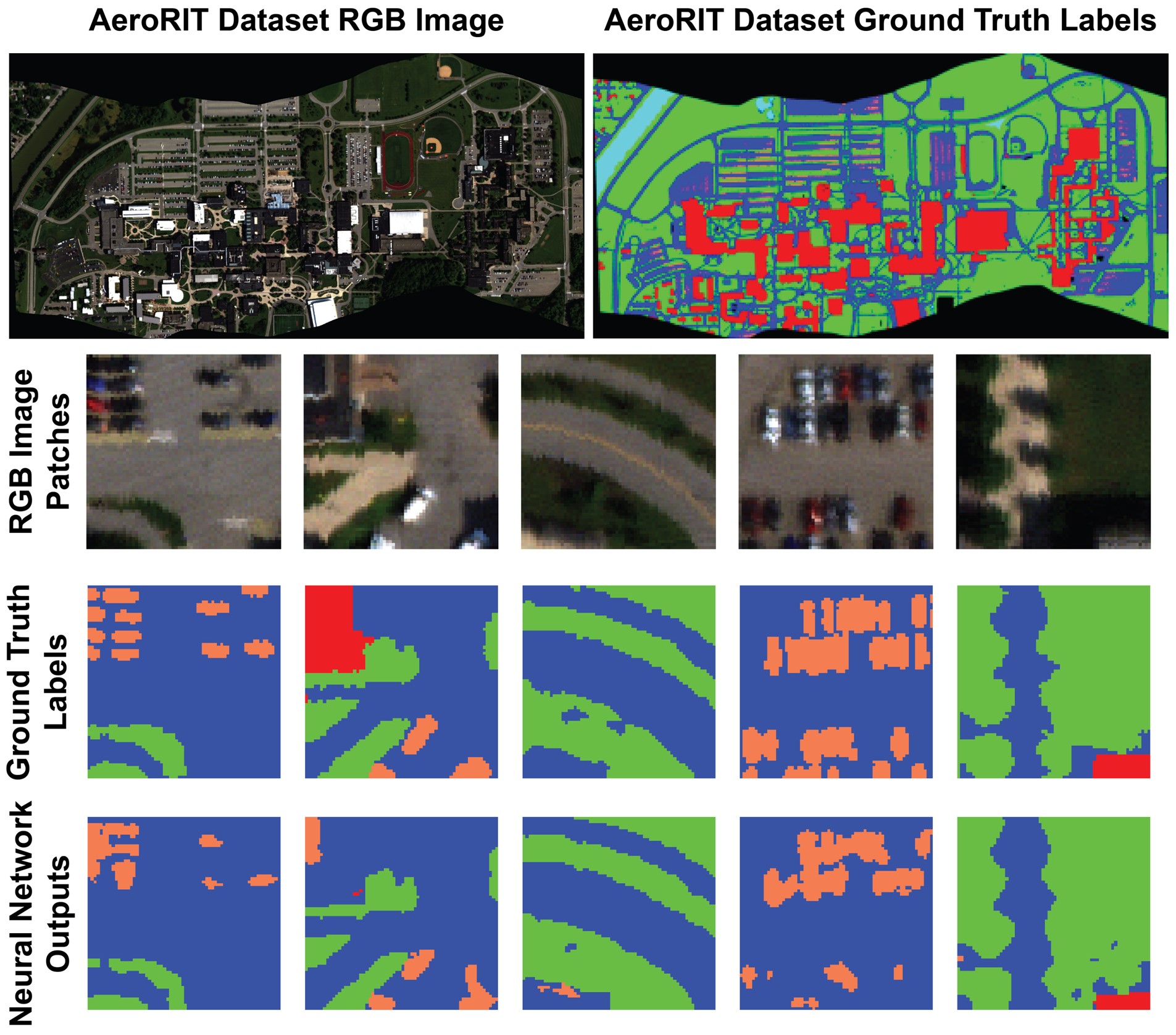
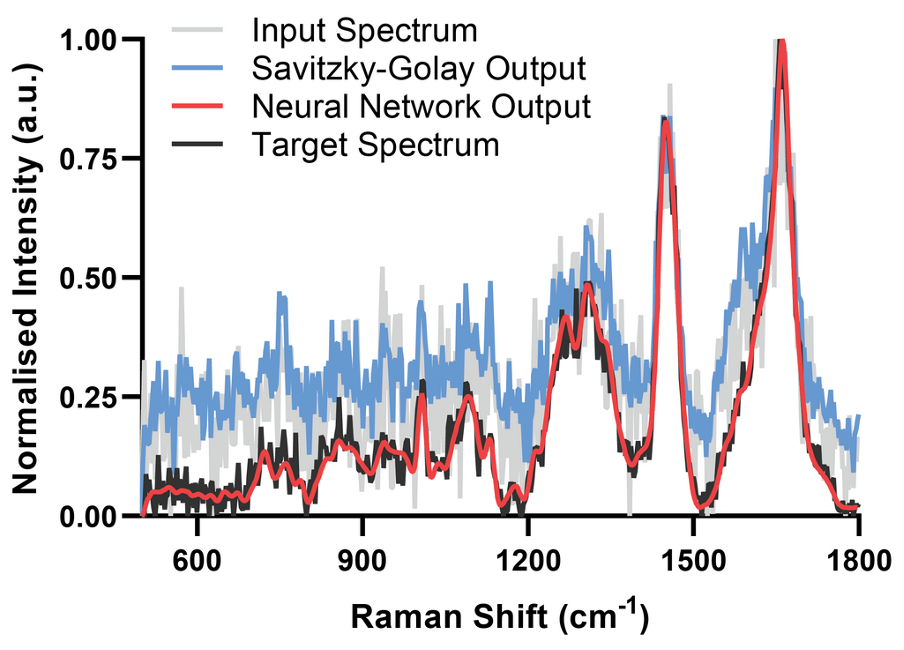
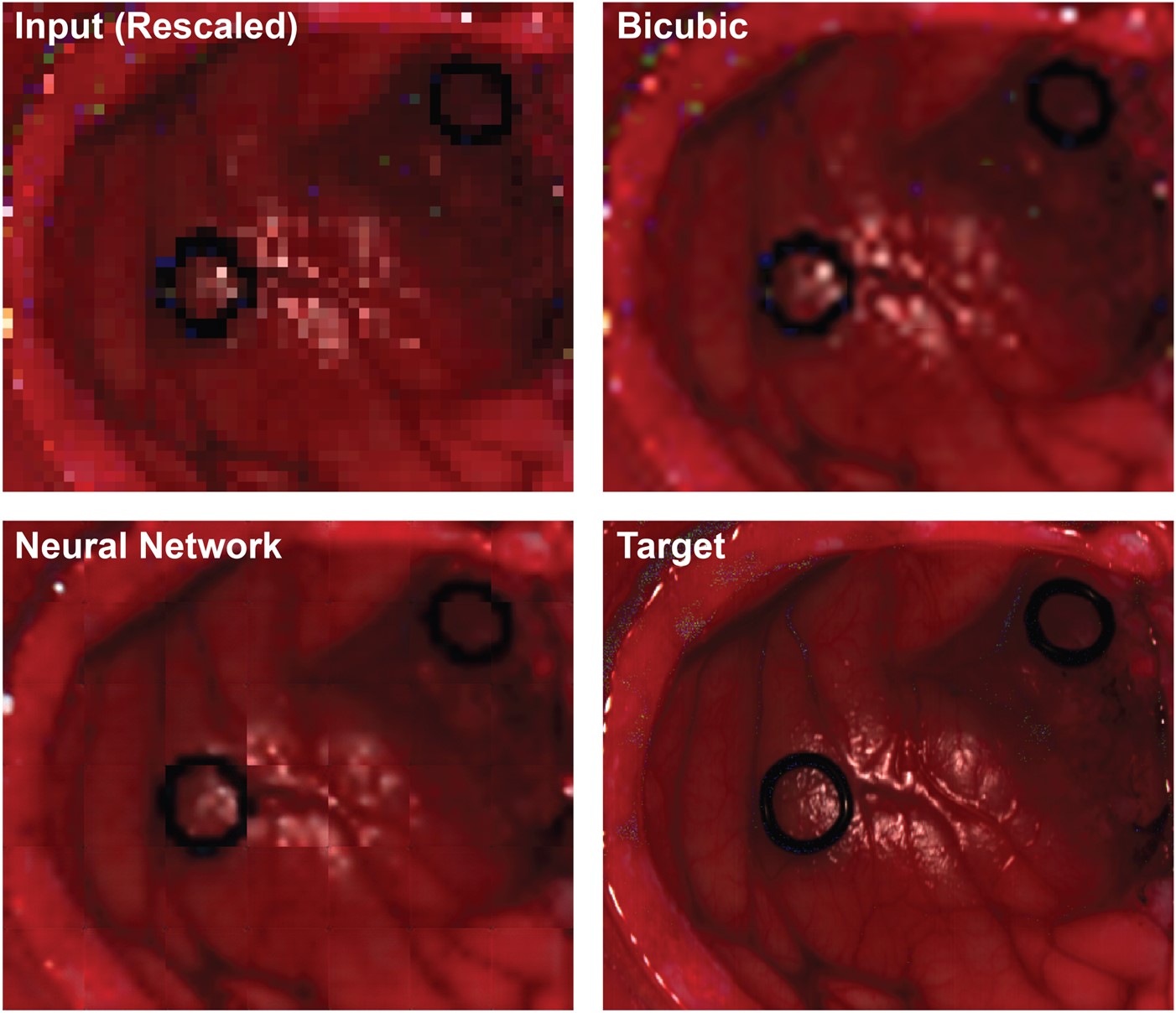
SpectrAI is a open-source framework bringing state-of-the-art AI to spectroscopy and spectral imaging from denoising to hyperspectral segmentation.